NXtec D001 Hereditary Cancer Panel 1
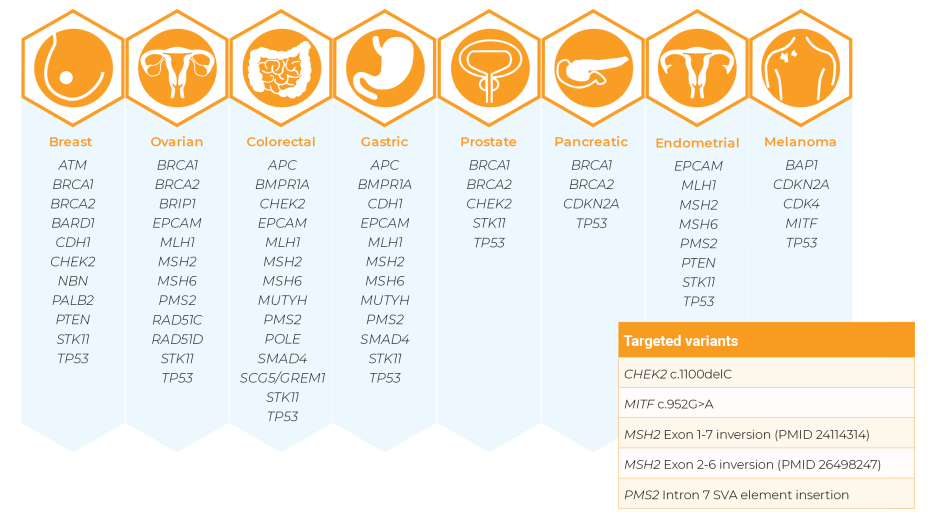
NXtec D001 Hereditary Cancer Panel 1 is a digitalMLPA assay that detects copy number variations in genes associated with hereditary predisposition to breast, ovarian, colorectal, gastric, prostate, pancreatic or endometrial cancer, or melanoma.

Contents: 723 probes, including 575 probes for 28 hereditary cancer-related genes (of which 3 probes detect the wild type allele of mutations in PMS2, POLE and BRCA2), and 7 mutation-specific probes for MSH2, MITF, PMS2 and CHEK2.

Tissue: genomic DNA isolated from human peripheral whole blood.

Application: research on predisposition to 8 common cancer types.

For research use only (RUO). Not for use in diagnostics.