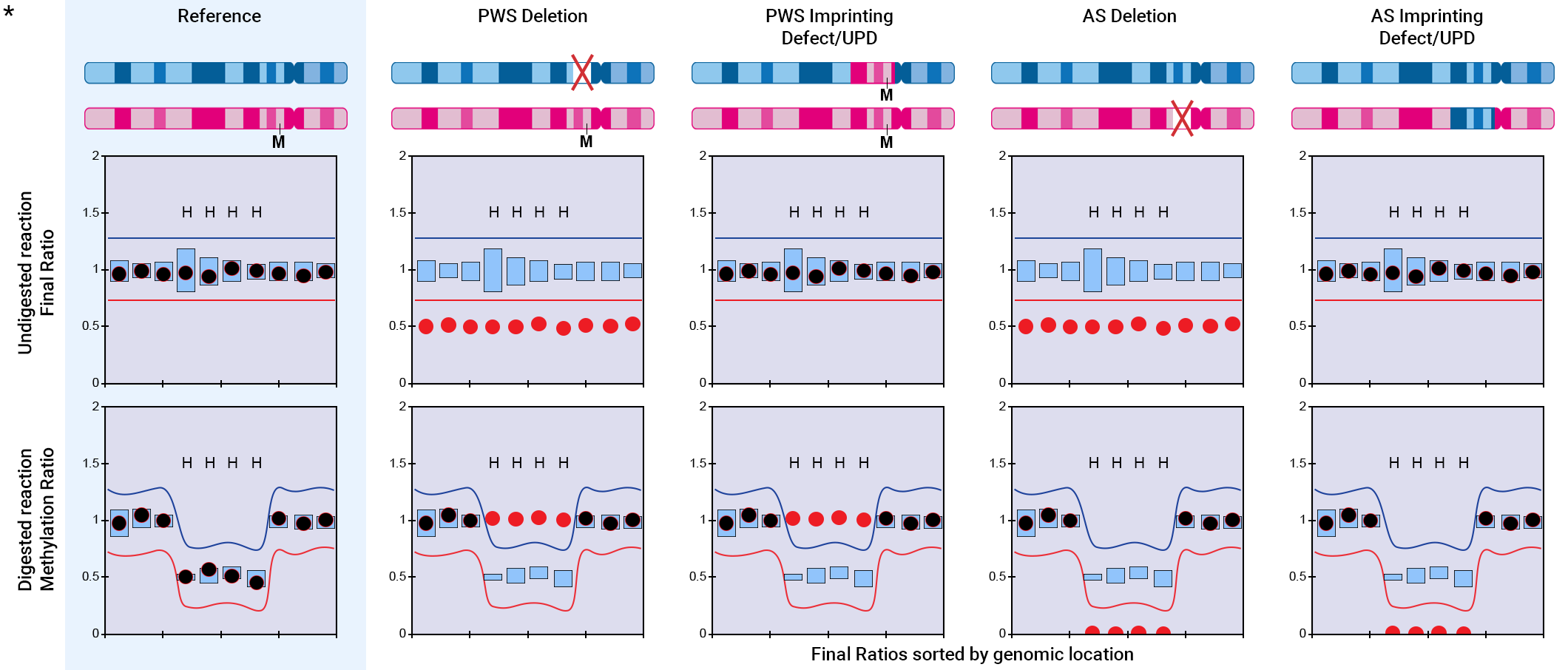
The figure below shows theoretical final ratios (from the undigested reaction) and methylation ratios (from the digested reaction) for the 15q11 region in cases of deletions or imprinting defects/uniparental disomy (UPD) associated with Prader-Willi syndrome (PWS) and Angelman syndrome (AS).
In cases of a paternal deletion causing PWS, the copy number of the target region is reduced to 1. Consequently, the final ratios decrease, and methylation ratios for methylation-specific probes (denoted by "H") increase, indicating full methylation in the remaining copy.
In contrast, for a maternal deletion leading to AS, the methylation ratios of methylation-specific probes (H) decrease, reflecting the absence of methylation in the single, unmethylated paternal copy of the target region.
For cases involving imprinting defects or UPD, no change in copy numbers occurs, thus final ratios remain unchanged. Only the methylation ratio is affected, depending on whether a paternal or maternal UPD or imprinting defect is present.
The table below provides an overview of the genomic and methylation patterns in the 15q11 region under different genetic conditions. It highlights variations in copy number, methylation percentages, and digestion ratios associated with Prader-Willi Syndrome (PWS), Angelman Syndrome (AS), and chromosomal duplications.
|
PWS Deletion |
PWS Disomy¥ |
Reference |
AS Disomy¥ |
AS Deletion |
Duplication |
| Genomic situation of the 15q11 region* |
M_ |
MM |
PM |
PP |
P_ |
PMM |
PPM |
| Copy number |
1 |
2 |
2 |
2 |
1 |
3 |
3 |
| Copy number ratio |
0.5 |
1 |
1 |
1 |
0.5 |
1.5 |
1.5 |
| % methylated |
100% |
100% |
50% |
0% |
0% |
70% |
30% |
| Methylation ratio after digestion |
1 |
1 |
0.5 |
0 |
0 |
0.7 |
0.3 |
* In this row, the paternal and maternal copies of the 15q11 region are indicated with a P or M, respectively.
¥ Next to uniparental disomy, PWS/AS can also be caused by aberrant methylation due to imprinting defects. With the ME028 probemix no discrimination between uniparental disomy and imprinting defects can be made.