Methylation-Specific MLPA
Methylation-specific MLPA (MS-MLPA) is a variant of the SALSA MLPA technique. With MS-MLPA, it is possible to semi-quantitatively profile methylation for multiple targets simultaneously.
MS-MLPA probes
Like regular MLPA probemixes, MS-MLPA probemixes contain probes that target a specific genomic sequence. Each probe consists of two parts: a left and a right probe oligonucleotide (LPO and RPO). Some of the probes in an MS-MLPA probemix target a region that contains a restriction site for the methylation sensitive HhaI endonuclease. These probes can be used for methylation profiling.
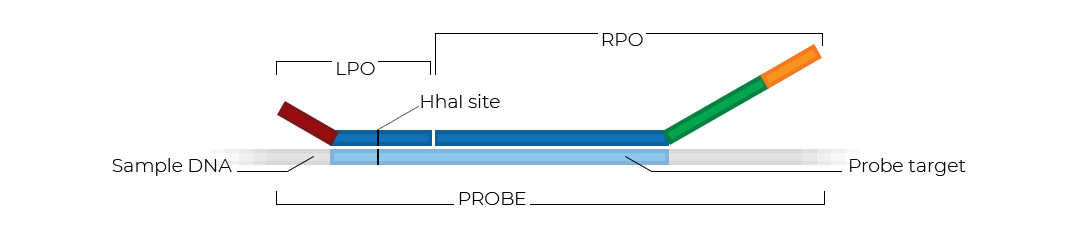
Principle of MS-MLPA
The probes hybridise to their target sequences on the sample DNA. The HhaI enzyme is then used to digest unmethylated double-stranded probe–DNA complexes.
Complexes with unmethylated sample DNA are digested. Digested probes cannot be amplified exponentially, and therefore produce no signal. Complexes with methylated sample DNA are not digested by HhaI, and will generate a normal signal.
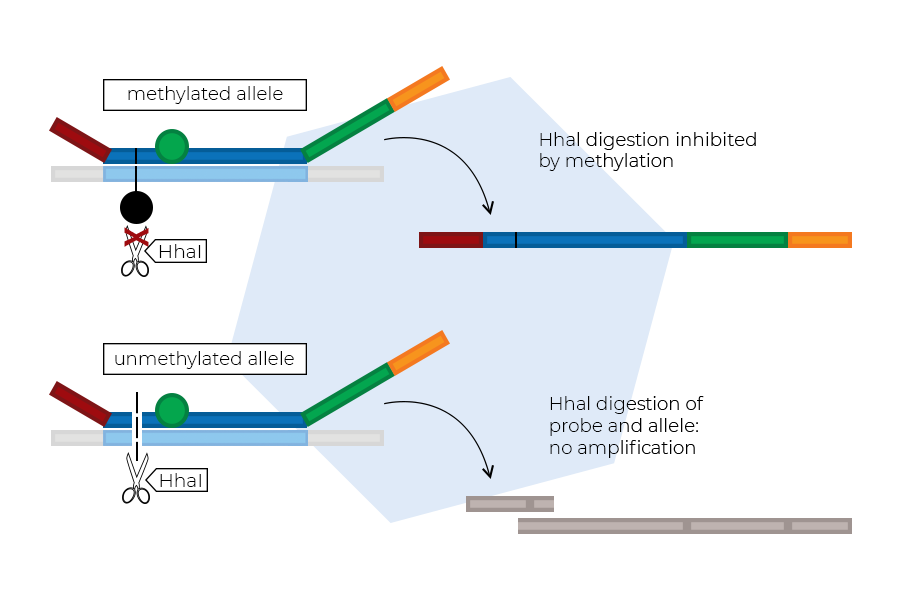
MS-MLPA reaction
The steps for MS-MLPA are highly similar to those outlined for MLPA. The most important difference is that the reaction is split into an undigested and a digested reaction after overnight hybridisation.
The undigested reaction is used for copy number determination. In the digested reaction, which is used for methylation profiling, the HhaI enzyme is added at the same time as the Ligase-65 enzyme.
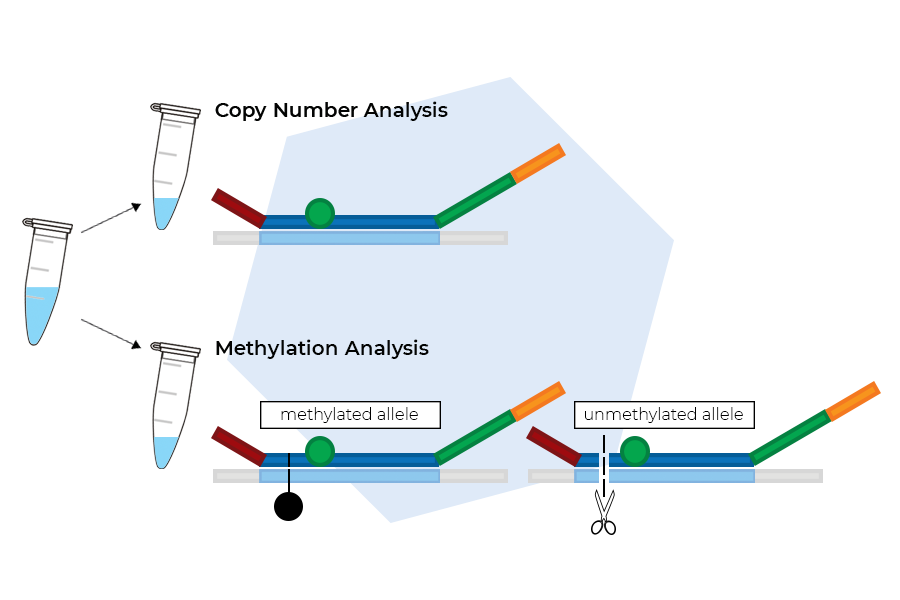
Data analysis
Coffalyser.Net is used to analyse the data. Copy numbers are determined using the undigested reactions in the same way as for regular MLPA. The methylation percentage in the DNA sample is determined by comparing each probe signal in the digested reaction with the corresponding signal in the undigested reaction.
More Information
Read our MS-MLPA General Protocol.
The MS-MLPA technique was first described in Nygren et al. 2005.