Principle of digitalMLPA
The principle of digitalMLPA is explained in the video below. Read on for a textual explanation. For alternative ways to view the video, see this support article.
digitalMLPA probes
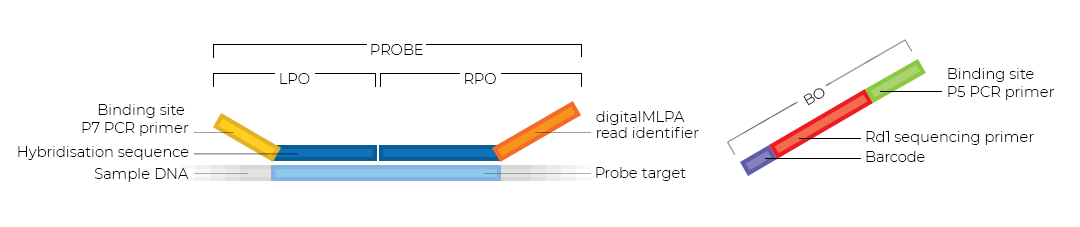
digitalMLPA probemixes have probes that target a specific genomic sequence. A digitalMLPA probe consists of two parts: a left and a right probe oligonucleotide (LPO and RPO). The LPO contains a P7 PCR primer and a DNA hybridisation sequence. The RPO contains a sequence for distinct digitalMLPA read identification and a DNA hybridisation sequence. A Barcode Oligo (BO) is added to every reaction. The BO contains a unique sequence for sample identification, along with Rd1 and P5 primer sequences.
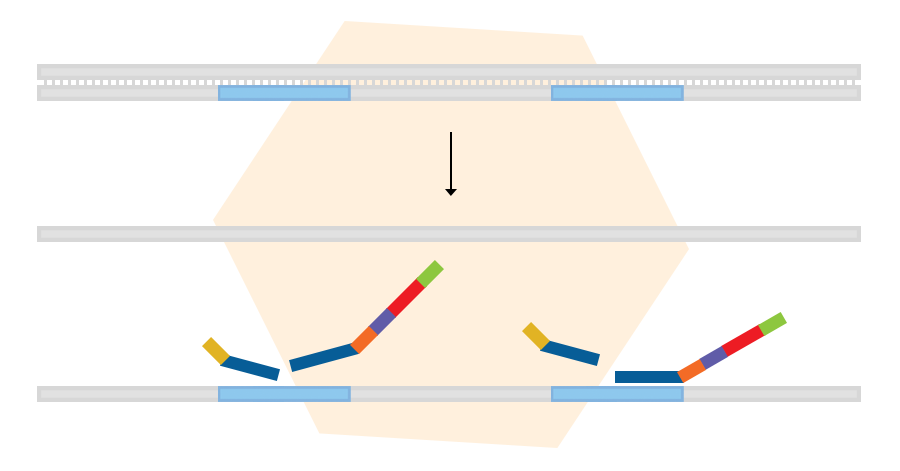
Sample denaturation and probe hybridisation
In the first step, purified sample DNA is denatured. This is followed by overnight incubation with digitalMLPA probe oligos. The LPO and RPO parts of each probe hybridise to immediately adjacent target DNA sequences.
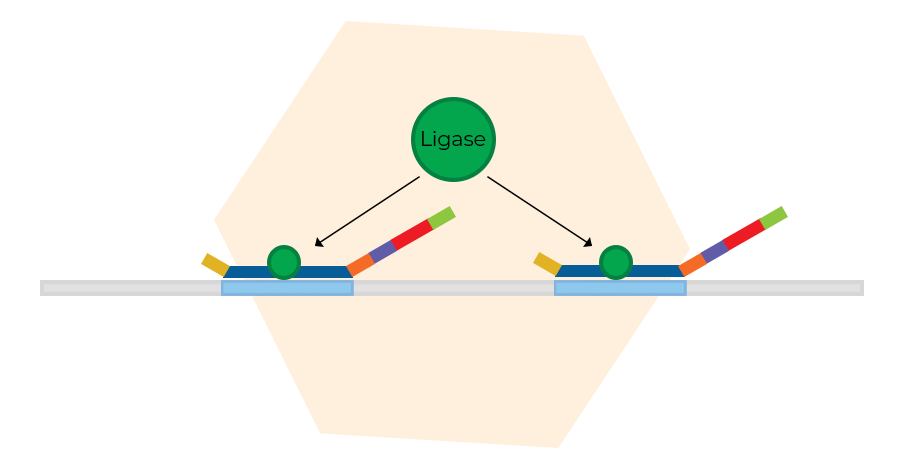
Probe ligation
The next step is the ligation of probe oligos that hybridised to immediately adjacent target sequences. No mismatches around the ligation site are permitted, making the ligation reaction highly specific. The number of probe ligation products is a measure for the number of target sequences in the sample. With digitalMLPA, it is possible to detect the number of ligation events.
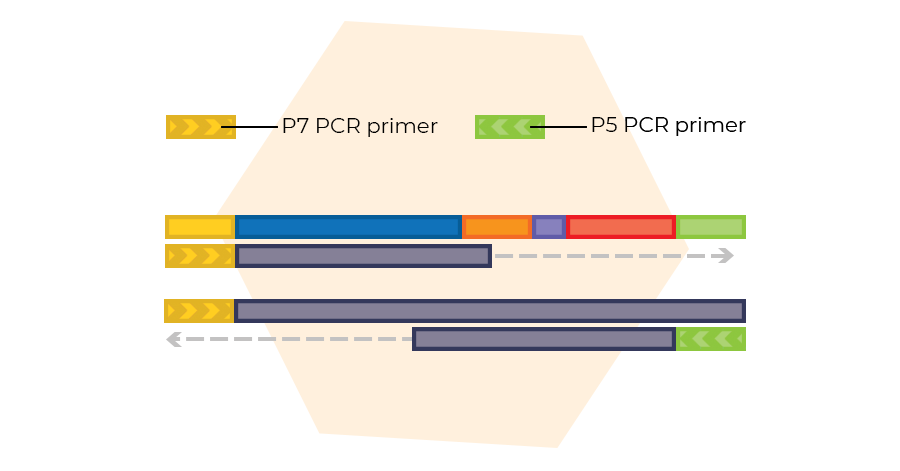
Probe amplification
Ligated probes are amplified in a multiplex PCR using a single P7/P5 primer pair. Only ligated probes are exponentially amplified, making the removal of unbound and non-ligated probes unnecessary.
Amplicon sequencing
PCR products are mixed in equal amounts, diluted, denatured, and loaded onto an Illumina sequencer. For each probe in all reactions, at least 600 single reads should be generated for reliable quantification.
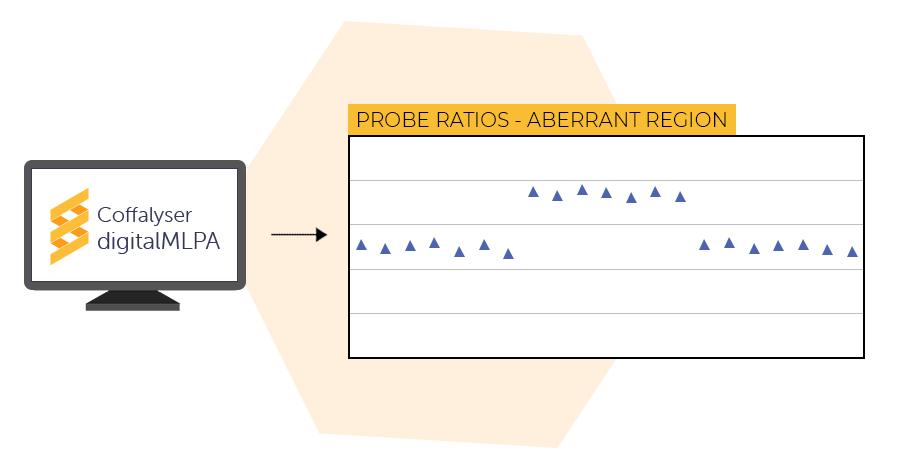
Data analysis
The final step is data analysis by Coffalyser digitalMLPA. Coffalyser digitalMLPA generates two easy-to-interpret reports for each sample in your experiment: one with general details and one with sample results. These reports indicate if sample and reaction quality were sufficient and highlight detected aberrant regions.
More Information
Read our digitalMLPA General Protocol.