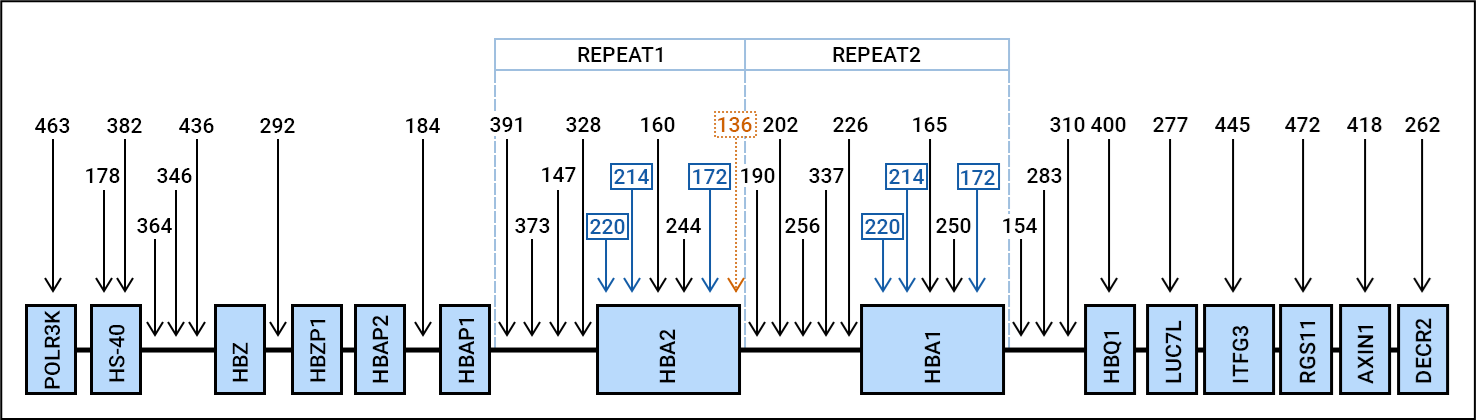
Alpha-thalassaemia is the most common inherited haemoglobin disorder in the world. It is characterised by a reduced production of the alpha-globin chain, resulting in a decrease in the total amount of haemoglobin. In normal adult life, about 97% of the total haemoglobin level comprises haemoglobin A (HbA), which is composed of two alpha- and two beta-globin chains. The remaining 3% of adult haemoglobin consists of HbA2 and HbF (foetal haemoglobin), consisting of two alpha chains combined with two delta-globin chains or two gamma-globin chains, respectively.
The alpha-globin chains are encoded by the haemoglobin alpha 1 (HBA1) and alpha 2 (HBA2) genes, located in the alpha-globin gene cluster on chromosome 16p13.3. Defects in the HBA genes can lead to two clinically significant forms of alpha-thalassaemia. In the lethal Hb Bart’s hydrops foetalis syndrome, the two HBA1 and two HBA2 copies are all absent or defect. In HbH disease, only one functional HBA copy remains. In addition, two alpha-thalassaemia carrier states can be discriminated: in alpha-thalassaemia trait (heterozygous α0-thalassaemia or homozygous α +-thalassaemia), two functional HBA copies remain, whereas in “silent” alpha-thalassaemia (heterozygous α +-thalassaemia), three functional HBA copies are present. Next to defects in the HBA genes, alpha-thalassaemia can also be caused by deletions in the upstream hypersensitive (HS) sites, which constitute the regulatory elements of the alpha-globin gene cluster.
Alpha-thalassaemia patients can present with a wide variety of clinical symptoms, ranging from very mild anaemia to severe transfusion-dependent haemolytic anaemia. The phenotype depends on the gene that harbours the mutation (HBA1 or HBA2), the type of mutation, and the number of affected alpha-globin genes.
Alpha-thalassaemia is inherited in an autosomal recessive manner, with about 85% of all alpha-thalassaemia phenotypes caused by genomic deletions in the HBA1 and HBA2 genes. Most of these deletions can be detected by the MLPA technique, including commonly occurring deletions such as the 3.7 kb deletion (-α3.7), the 4.2 kb deletion (-α4.2), the South-East Asian deletion (--SEA), and the Filipinian deletion (--FIL). The remaining 15% of the alpha-thalassaemia cases result from one of at least 70 different point mutations, usually located within the HBA2 gene (Higgs and Weatherall, 2009; Harteveld and Higgs, 2010). The most common non-deletion mutation, which is frequently seen in Southeast Asia, is Hb Constant Spring, resulting from a mutation in the stop codon of the HBA2 gene. This mutation leads to the production of an elongated α-globin chain. Hb Constant Spring is produced in very small amounts because its mRNA is unstable. Heterozygotes for elongated globin chain variants such as Hb Constant Spring present with an α0-thalassemia phenotype. Presence of the Hb Constant Spring mutation can be detected by the P140 probemix.
In addition to many deletion types, several duplications have also been described in the alpha-globin gene cluster. These duplications vary in size, ranging from only a single duplicated HBA gene to large segmental duplications of the complete alpha-globin gene cluster, including the regulatory elements. Duplication of one or both HBA genes is clinically benign. However, when co-inherited with a beta-thalassaemia mutation, an HBA duplication leads to a more severe phenotype in beta-thalassaemia patients because it aggravates the balance between alpha- and beta-globin chains.
More information on alpha-thalassaemia is available on http://www.ncbi.nlm.nih.gov/books/NBK1435/.